Research concept
Jump to Research projects, Using microorganisms, Techniqus, Equipments
Microorganisms have specific functions in different environments
Microorganisms in a given environment have specific functions in response to environmental factors such as temperature and pH, as well as essential growth factors such as carbon and nitrogen sources. This function is ultimately derived from the genes encoded in its chromosome of the cells. Our goal is to discover and reveal the unique functions of various microorganisms.
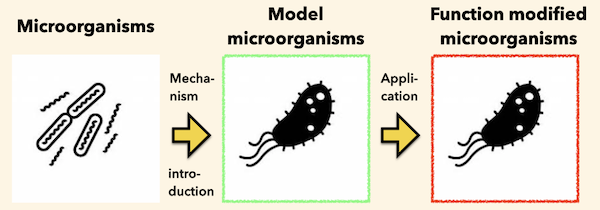
Analyzing Microbial Functions in Microbial Cells
Most microbial research to date has involved direct analysis or modification of specific microorganisms. Although much knowledge has been obtained by such methods, in most cases, the unique functions and mechanisms of a particular microorganism can only be utilized in its own cells. However, advances in genetic engineering have enabled large-scale genome modification of some microorganisms, making it possible to introduce and reproduce the functions (principles) of specific microorganisms. In our laboratory, we aim to understand and apply the specific function of a particular microorganism and to reproduce that function in a different microorganism. In addition, some genes are difficult to functionally express in model microorganisms. Until now, genes that do not show such functions have been left as they are, and no progress has been made in accumulating knowledge on why these genes do not function in different microorganisms. Therefore, we aim to accumulate such knowledge and technologies for improvement.
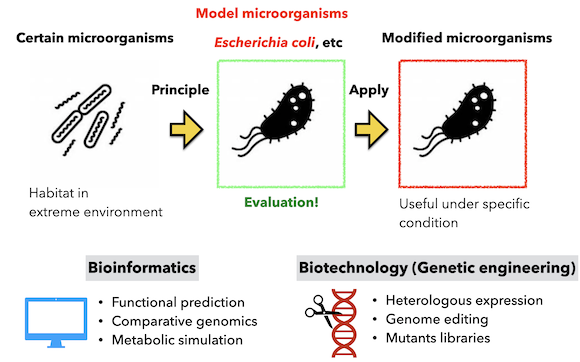
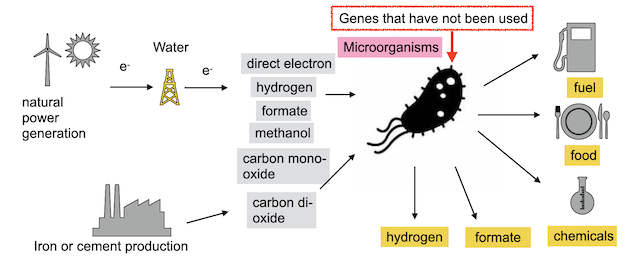
Development of microorganisms that applied unused genes to utilize environmental energy
By conducting functional analysis using microorganisms, the genes are already functioning in the cells. By increasing the number of genes that function in cells and utilizing them in this way, we are working on the development of microorganisms that will make it possible to manufacture products and produce energy alone based on environmental energy using microorganisms.
Research projects
-
Research on the unique metabolic mechanisms of microorganisms
Propionate oxidation under anaerobic conditions requires a syntrophic relationship between propionate-oxidizing bacteria and hydrogenotrophic methanogens, but the oxidation of propionate involves unique metabolic mechanisms of each microorganism. Our research focuses on the mechanism by which propionate oxidizing bacteria produce hydrogen and formic acid from propionate. On the other hand, genes of microorganisms living in extreme environments may be able to confer their unique metabolism to other microorganisms. Currently, we are attempting to express genes unique to archaea living in highly alkaline environments in Escherichia coli. -
Research on metabolism and robustness of microorganisms
We are studying the robustness of microorganisms, especially their response to heat and its relationship to metabolism. We are investigating which pathways are required by E. coli for growth at high temperatures and how the intermediate metabolites synthesized by these pathways are required for growth at high temperatures. On the other hand, we are studying how to optimally construct metabolic pathways in E. coli that produce specific substances. -
Research on visualization of gene function using microbial cells
Genome modification is able to be carried out process, and fluorescent proteins and tags are helpful to monitor the expression of target genes. Using these techniques, we are investigating methods to visualize internal metabolism and protein expression, especially in E. coli.
Using microorganisms
- Escherichia coli
- Pelotomaculum thermopropionicum
- Methanothermobacter sp.
- Bacillus subtilis
- Corynebacterium glutamicum
- Cupriavidus necator
- Shewanella oneidensis
Techniqus
- Microorganism Culture
- Culture medium
- Culture with Oxygen
- Culture without Oxygen
- Culture with hydrogen
- Ultra-low temperature preservation of microorganisms
- Observation of bacteria
- Fluorescence microscopy
- Fluorescence observation on plates
- DNA extraction
- Genome extraction
- Plasmid extraction
- Agarose gel electrophoresis
- DNA processing
- PCR
- Gibson assembly
- iVec3
- Microbial transformation
- Heat shock
- Electroporation
- Conjugation
- P1 transduction
- lambda RED recombination
- Protein extraction and analysis
- Cell disruption
- Ultracentrifugation
- SDS-PAGE
- Western blotting
- Protein purification
- Substance analysis
- Spectroscopic analysis
- Fluorescence analysis
- Gas chromatography
- Liquid chromatography
- Mass spectrometry
- Bioinformatics
- Database Search
- PCR Primer Design
- Gene Function prediction
- Genome Information Analysis
- Transcriptome analysis
- Other Omics Analyses
- Metabolism Simulation
- Flax Balance Analysis
- Programming languages and programs
Equipments
- Balances
- pH Meter
- Pure Water Production Equipment
- ADVANTEC
- Sterilizer
- Clean bench
- Constant temperature machine
- Shaking temperature controller
- TAITEC
- Eppendorf ThermoMixer C
- Anaerobic chamber
- Bacbasic
- Cell disruption system
- MPB FastPrep 24
- French press (common)
- Centrifuge
- KUBOTA 3740
- KUBOTA PlateSpin3
- Hitachi Ultracentrifuge (common)
- Thermal Cycler
- Applied Biosystems GeneAmp PCR System 2700
- Bio-rad T100
- Takara
- Electrophoresis system
- Takara Mupid-exU
- Transilluminator
- UV
- LED 470 530
- Image Analyzers
- ATTO PrintGraph
- Spectrophotometer
- Shimadzu UV-1850
- Berthold biochrom Libra S12
- Fluorescence Spectrometer
- JASCO FP-6200
- Fluorescence microscope
- Nikon type 120
- Bioanalyzer
- Agilent 2100
- Deep freezer
- Freezer
- Refrigerator
- Refrigeration Chamber
- Gas Chromatograph
- Shimadzu GC-8A TCD
- Liquid Chromatograph
- Waters Alliance (common)
- Shimadzu LDx (common)
- Protein Chromatograph
- Cytiva AKTAprime plus (loaner)